2021 Bruins-In-Genomics Summer Undergraduate Research Program
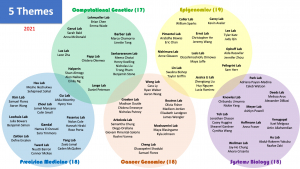
2021 B.I.G. Summer Participants
| Lab PIs | Mentors | Students |
|---|---|---|
| PATRICK ALLARD | Ramanuj Sarkar | |
| Medhini Sosale | ||
| VALERIE ARBOLEDA | Sarah Spendlove | Desiree Cervantes |
| Sarah Spendlove | Matthew Moleres | |
| PAUL BOUTROS | Taka Yamaguchi | Anna Neiman-Golden |
| Chenghao Zhu | Siddharth Mahesh | |
| Julie Livingstone | Karthik Guruvayurappa | |
| Chenghoa Zhu | Ali Mobedi | |
| Taka Yamaguchi | Philippa Steinberg | |
| MANISH BUTTE | Audrey Yang | |
| JEFF CHIANG | Johnson Chen | Asiya Dalak |
| Johnson Chen | Eleanor Carr | |
| Akos Rudas | Simon Lee | |
| JASON ERNST | Ha Vu | Aahna Rathod |
| Ha Vu | Saiyang Liu | |
| Ha Vu | Elijah Jones | |
| NANDITA GARUD | Ricky Wolff | Kaysa Pfannmuller |
| Mariana Harris | Julianna Perez | |
| Xiaolu Guo | Jianche Liu | |
| WILLIAM HSU | Ruiwen (Rina) Ding | Garrett Duncan |
| JIMMY HU | Leah Ye | Hanifa Maswali |
| STEVEN JACOBSEN | Zhenhui Zhong | Karisa Ke |
| COLIN KREMER | Aarshi Jain | |
| HONGHU LIU | Jie Shen | Talia Villa |
| Jie Shen | Fabely Moreno | |
| KIRK LOHMUELLER | Saige Daines | |
| CHONGYUAN LUO | Matthew Heffel | Catalina Holguin |
| Matthew Heffel | Maya Brawer-Cohen | |
| AARON MEYER | Jackson Chin | Carly Gordon |
| Jackson Chin | Jalissa Emmens | |
| ALIREZA MOSHAVERINIA | Isabela Molina | |
| Mary Torres Guzman | ||
| ROEL OPHOFF | Marcelo Francia | Ethan Concepcion |
| Marcelo Francia | Zachary Jordan | |
| PAIVI PAJUKANTA | Asha Kar | |
| MATTEO PELLEGRINI | Sabrina Perez | |
| Ashley Lopez | ||
| Wenbin Guo | Junxi Feng | |
| Wenbin Guo | Hongxiang Fu | |
| Salvador Ayala | ||
| TAMER SALLAM | Zhengyi Zhang | Vivien Su |
| SRIRAM SANKARARAMAN | Noela Wheeler | |
| Aakarsh Anand | ||
| Prateek Anand | ||
| Michelle Johnson | ||
| MARC SUCHARD | Fan Bu | Keyi Xue |
| HUNG TON-THAT | Amanda Fuenzalida | |
| DANIEL TWARD | Bryson Gray | Osagie Aimiuwu |
| Bryson Gray | Tiffanie Crumbie | |
| Bryson Gray | Sarthak Tiwari | |
| SHARMILA VENUGOPAL | Daniel Felipe Forero | |
| Fiona Latifi | ||
| Haoxuan (Harry) Zhang | ||
| Cameron Gill | ||
| CLAUDIO VILLANUEVA | Mirian Krystel De Siqueira | Victoria Pellot |
| CUN-YU WANG | Jing Wang | Kennedey Boyette |
| Jing Wang | Eduardo Del Rio | |
| JENNIFER WILSON | Emily Liu | |
| Jarod Le | ||
| DAVID WONG | Karolina Kaczor-Urbanowicz | Rachelle Chanthavong |
| Misagh Kordi | Timothy Lindsey | |
| Misagh Kordi | Manuel Cortes | |
| GRACE XIAO | Jonatan Hervoso | Rohan Chatterjee |
| Jonatan Hervoso | Raymond Benitez | |
| XIA YANG | Monty Blencowe | Nuoya Jiang |
| Monty Blencowe | Imri Haggin | |
| Monty Blencowe | Darren Wijaya | |
| Monty Blencowe | Ning Wang | |
| JASMINE ZHOU | Ran Hu | Tony Zhang |
| Ran Hu | Harinarayana Mellacheruvu |
2021 B.I.G. Summer Poster Abstracts
ALLAHVERDIAN: Constructing a Novel Protein-Protein Interactome for Neuroimmune Modulation of Neural Excitability
ARTIN ALLAHVERDIAN (1,3), Roxane Knorr (2,3), Jakob von Morgenland (3), Sharmila Venugopal (1,2,3)
- BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
- STEP UP Scholar Program, National Institute of Health
- Neural Dynamics Group, The Department of Integrative Biology and Physiology, UCLA
Neuroinflammatory cytokines mediate acute and chronic roles in moderating nervous system disease states. However, knowledge of their dynamic interactions with ion channel proteins (ICPs) and other biomolecules important for neural functions is sparse, largely owing to the complexity and scale of such interactions. To address this, we present the development of a protein-protein interactome (PPI). Beginning with systematic literature curation and meta-data generation, I will discuss our novel functional interaction score (FIS) to quantitate cytokine modulation of neural excitability function and its mediators, the ICPs. Using the FIS as the basis for edge weights, I will further discuss generation of a PPI using Cytoscape. As the future direction, we will construct a comprehensive PPI network to identify the hub proteins that may play a crucial role in neurodegenerative disease progression.
ALMOGY, NAIBERG, NG: Leveraging methylation risk scores to account for unmeasured confounding in epigenome-wide association studies
SIVAN ALMOGY1, EMILY NG1, ALON NAIBERG1, Michael Thompson2, Eran Halperin3
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Bioinformatics Interdepartmental PhD Program, UCLA
3 Department of Computational Medicine, UCLA
DNA methylation levels in the genome evolve to reflect environmental and lifestyle factors, the use of medication, and the onset of disease. As many factors affect DNA methylation levels, epigenome-wide associations are particularly at risk for confounding. In this study, epigenetic data of rheumatoid arthritis cases and controls were re-assessed to find CpG sites that are associated with the disease using an epigenome-wide association study (EWAS), while accounting for medications and other unmeasured confounders using externally developed methylation risk scores (MRS). After controlling for the imputed confounders, several associations no longer showed significant association signal –in particular an association at a locus responsible for T cell function was not significant after accounting for intake of immunosuppressive drugs. Given the results of this study, controlling for imputed medications and other confounders in future studies may provide useful insights into the CpG sites associated with other diseases.
AROS, DIBIASI: Implementing layer structures to assess a neural network’s function as a nonlinear dimensionality technique
MELISSA AROS1, ALEXANDER DIBIASI1, Melanie Tu2, Serena Hughes3, Eric J Deeds4
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Deeds Lab Staff, UCLA
3 Bioinformatics Interdepartmental PhD Program, UCLA
4 Department of Integrative Biology and Physiology, UCLA
Researchers reduce the dimensionality of sc-RNA sequencing datasets before performing downstream analysis because the data is so large. However, Dr. Deeds’ lab discovered that existing techniques to do this introduce high distortion to the data. The lab is developing a neural network-based dimensionality reduction method called the DeepEmbedder which will minimize distortion. There is a metric known as Jaccard Distance which can be used to quantify distortion, where a low Average Jaccard Distance (AJD) indicates low distortion. Our project focused on analyzing how the layer structures used in the DeepEmbedder affect the final AJD of the embedding. We found the structure 3-100-50-2 had the lowest AJDs; whereas, the structure 3-4-5-6-7-8-9-2 had the highest AJDs. Our contribution to this project is showing that altering the layer structure does impact the resulting embedding, a discovery that Dr. Deeds’ lab will use to further improve the DeepEmbedder.
AVELAR: Differential Regulation by AP-1 Genomically linked to Oncogenesis in Non-Small Cell Lung Cancer
KEVIN AVELAR DIAZ1 2, Xianglong Tan2, Alisha Flora2, Michael Kronenberg2, Michael Carey2
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Biological Chemistry, David Geffen School of Medicine, UCLA
Expression of transcription factors (TFs) is subject to control by enhancers. Abnormalities in promoter-enhancer interactions are associated with differential TF levels linked to oncogenesis. However, it remains difficult to identify oncogenic TFs involved in the development of specific cancer types. To discover cancer-specific TFs, we identified overexpressed TFs in cancer using RNA-seq, and developed a methodology to integrate promoter-capture Hi-C and multi-omics data with motif analysis. This method was applied to non-small cell lung cancer (NSCLC) cells and normal lung bronchial epithelial cells. We found significant enrichment of the AP-1 TF family in enhancers of overexpressed genes in NSCLC cells. ChIP-seq analysis confirmed our findings, as NSCLC cells had greater genome-wide binding of AP-1 TFs at promoters, enhancers, and super-enhancers. These findings suggest the AP-1 family is involved in transcriptional regulatory changes in NSCLC development. The methodology implemented here could potentially be applied to identify oncogenic TFs in other cancers.
BALD, MCDONALD: Differences in Genetic Diversity in the Gut Microbiome Between Hosts with Westernized and Non-Westernized Diets
SARAH BALD1, ANNA L. MCDONALD1, William R. Shoemaker2, Nandita R. Garud2,3
1BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2Department of Ecology and Evolutionary Biology, UCLA
3Department of Human Genetics, UCLA
The human gut microbiome is a dynamic ecosystem that plays a crucial role in human health and is ecologically and evolutionarily shaped by the behavior of its host. A major host behavior is diet, as non-westernized hosts have diets rich in fiber and complex carbohydrates, while Westernized hosts consume more processed foods. Because microbes require additional pathways to metabolize complex molecules, it is likely that non-Westernized hosts harbor higher levels of genetic variation as well as gene content and function, but few analyses have been performed to test this hypothesis. In this study, we compared the nucleotide diversity from abundant microbial species between 163 non-Western hosts from Africa and 180 Western hosts from North America. We found that the genetic diversity of genes that were present in all hosts was higher among African hosts, that African hosts harbored many unique genes, and that the disproportionately enriched genes within a particular group encoded metabolic pathways known to effect host health. Our results show that host diet has shaped the genetic diversity and composition of microorganisms within the human gut.
BARRON, MCKEE: The impact of clinical-grade versus research-grade image quality on volumetric analysis of T1 MRI brain scans
NUUTTI BARRON1, CONNOR MCKEE2, Daniel Tward3
- BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
- BIG Summer Program, Institute for Quantitative and Computational Biosciences, UT Austin
- Department of Computational Medicine, Department of Neurology, UCLA
Neurodegenerative diseases are a critical problem facing our aging population. Disease monitoring can be enhanced by measuring the volumes of neuroanatomical structures, performed noninvasively through MRI. Computational approaches for this have been validated in high-grade research scans, but not in low-grade clinical scans. Our goal is to determine how much accuracy is reduced in clinical-grade images. We used 18 research-grade T1 weighted MR images, with 287 structures annotated. We simulated clinical-grade images by degrading these with noise, blur, and anisotropic resolution. We calculated deformable transformations to align “atlas images” to degraded “clinical images” for each pair, under 28 conditions. We performed volumetric analysis on the aligned atlas, and reported accuracy for cortical and subcortical gray matter, white matter, and others. These results provide quantitative measures of accuracy when performing volumetric analysis on clinical-grade images. This is an important step toward analyzing electronic health records to study neurodegeneration at large scale.
BISHOP: Mechanisms associated with diverse tumor phenotypical changes after cell-cell fusion
SWALINA BISHOP1, Mengtao Li2, Lin Yi-Ling2 ,
1 Big Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Division of Diagnostic and Surgical Sciences, UCLA School of Dentistry
SCCVII/SF tumor cell lines were co-cultured with mouse bone marrow macrophages. Used as a fusion agent, RANKL was added to the culture media to induce tumor-macrophage fusion. After being injected into the mice, extreme growth patterns, large vs. little/no growth were noted among the SCCVII/SF-macrophage clones. Also, flow cytometry was used to quantify the contents in the tumor to understand how TAM and T-reg cells are associated with the tumor growths Since the cause for the divergent tumor growth is unknown, we performed RNAseq on the tumor clones and analyzed the RNA datasets for the control and the large tumors using FastQC, Trimmomatics, and hiSTAT2 for quality control and read alignment. The quality controls for all the raw reads were good and had excessive duplicated sequences. We used feature counts and Deseq2 for differential gene expression. Identifying the candidate genes could help to diagnose and treat cancer.
BISWAS, CHAN: End to End In Silico Framework for CRISPR Experiment SImulation and Testing
ARABDHA BISWAS1, ERIC CHAN1, Harold Pimentel2,3
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Computational Medicine, David Geffen School of Medicine, UCLA
3 Department of Human Genetics, David Geffen School of Medicine, UCLA
Highly parallel CRISPR screens have enabled dissection of putative regulatory regions in a single experiment, and new assays are constantly being invented. However, a major challenge of these experiments is determining design: which regions are probable? How many cells are required? How does inference behave in various conditions? One such screen tests for functional regions of a protein by modifying tiling windows of the gene’s putative functional regions. We developed a simulation framework that allows us to specify various experimental conditions and test the experiment’s accuracy. Our framework adapts an existing model used for simulating in vivo CRISPR screen experiments. Through this model, we are able to in silico create mutations, simulate effects, generate cell counts, and test which mutations are recoverable computationally.This framework allows for researchers to explore experimental configurations resulting in varying of specificity and sensitivity of CRISPR screens before performing an experiment, thus not wasting effort on non-recoverable regions.
BLASINGAME, JOHNSON: The Effect of the Physical Properties of Biomaterials in Regulating the Secretome of Dental Mesenchymal Stem Cells for Tissue Engineering
MAYA BLASINGAME1, KYLA JOHNSON1, Alireza Moshaverinia2,3,4, Sevda Pouraghaei Sevari2,3,4
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Laboratory of Biomaterials Innovation and Tissue Engineering (BITE), School of Dentistry, UCLA
3 Weintraub Center for Reconstructive Biotechnology, School of Dentistry, UCLA
4 Division of Advanced Prosthodontics, School of Dentistry, UCLA
Mesenchymal stem cells (MSCs) have become essential in tissue repair and regeneration via their abilities to differentiate as well as their paracrine functions that help aid in immunomodulatory responses and trophic regenerative factors. However, MSCs are limited in their effectiveness because of the immune system’s negative response. Thus, this project looked at the role of biomaterials and their properties in regulating the relationship between the immune response and MSCs in tissue engineering. This was done by analyzing the effectiveness of stiffness and porosity of biomaterials on gene expression and regulation of the secretome of MSCs. It was found that the physical properties of biomaterials, specifically, enhanced the effectiveness of MSCs in tissue engineering by physically protecting MSCs and upregulating several paracrine factors. The clinical application of MSCs combined with the aid of biomaterials is a promising new area of regenerative medicine because of their versatility to regulate various biological processes.
BOWERS, SIMON: Applying Latent Dirichlet Allocation for Dimensional Phenotyping of Clinical Notes in Serious Mental Illness
JULIA BOWERS1, BENJAMIN SIMON1, Juan De la Hoz2, Loes Olde Loohuis3
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Bioinformatics Interdepartmental PhD Program, UCLA
3 Department of Psychiatry and Biobehavioral Sciences, David Geffen School of Medicine, UCLA
Serious mental illnesses such as bipolar disorder (BD), schizophrenia (SZ), and major depressive disorder (MDD) are heterogeneous disorders with overlapping symptomatology. Trans-diagnostic and dimensional models, rather than traditional categorical ones, may provide insight into disease mechanisms. We used NLP to analyze over 40,000 intake notes from Colombian psychiatric electronic health records. Using Latent Dirichlet Allocation, we modeled notes as mixtures of “topics” representing relevant clinical dimensions and visualized them with t-SNE. Topics represented coherent groups of concepts such as ‘substance use’, ‘alcohol’, and ‘cannabis’ or ‘hallucinations’, ‘delusions’, and ‘orientation’. Topics were associated with diagnosis, gender, age, severity, and conversion from MDD to BD. Clustering patients with mental illness using electronic health record data may prove more useful than existing diagnostic categories in providing insights into disease mechanisms and progression, allowing for the development of individualized treatments.
CHACON: Identifying Conserved and Unique Bacterial Mutations for Combination Drug Resistance
JONATHAN CHACON1, Natalie Lozano-Huntelman2, Pamela Yeh2
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Ecology and Evolutionary Biology, UCLA
The diverse and dynamic repertoire of evolutionary maneuvers bacteria use to obtain resistance against antibiotics creates a major public health concern. Due to the increasing amount of multi-drug resistance, focus has been placed on repurposing current antibiotic treatments as combinational multi-drug treatments. However, the multifactorial nature of combinational drug therapies makes it challenging to assess how bacterial drug resistance will evolve. We use whole-genome sequencing to characterize antibiotic resistance genes in strains of Staphylococcus epidermidis that evolved resistance against a three-drug combination. We examined four three-drug combinations that consisted of piperacillin, tetracycline, and the third drug being either; chloramphenicol, doxycycline, erythromycin, or neomycin. Importantly, each three-drug combination surmises to a highly synergistic interaction that inhibits bacterial growth greater than expected from individual single-drug exposure. By identifying drug resistance genes at the whole genome level, we aim to characterize the evolutionary path that S. epidermidis takes to obtain combinational antibiotic resistance.
CHAMORRO, TANG: Testing marine microbial biogeography in the Coral Triangle
MARCO CHAMORRO1, LINETTE TANG1, Eric Caldera2, Paul Barber2
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Ecology and Evolutionary Biology, UCLA
Indonesia’s Coral Triangle is the most biodiverse marine ecosystem on Earth. While the diversity of metazoans is characterized by biogeographic species turnover, little is known about the dispersal of microorganisms. Baas Becking’s “Everything is everywhere but the environment selects” hypothesis posits that microbes lack biogeography and dispersal limits. Instead, microbial dispersal patterns may be shaped by symbiotic associations with metazoan hosts. We used data from Autonomous Reef Monitoring Structures deployed across the Coral Triangle and 16S metagenetic sequencing to determine whether reef bacteria have biogeographic patterning. Jaccard community distance among ARMS correlated with geographic distance, indicating dispersal limitations. Although recent interpretations of EEBES suggest that ubiquitous dispersal is determined by a 1mm size threshold, we found isolation by distance correlations to be strongest in ARMS size fractions below 1mm. Further research should integrate the impact of environmental factors such as pollution on microbial biogeography in the increasingly threatened Coral Triangle.
CHEN: Influence of Age Structure on the Inference of the Distribution of Fitness Effects
BRIAN CHEN1, Chris Kyriazis2, Kirk Lohmueller2,3,4
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Ecology and Evolutionary Biology, UCLA
3 Interdepartmental Program in Bioinformatics, UCLA
4 Department of Human Genetics, David Geffen School of Medicine, UCLA
The distribution of fitness effects (DFE) describes the effects of new mutations on fitness. Fitdadi is a recently developed software program that can infer the DFE using the site frequency spectrum (SFS), which describes the distribution of allele frequencies in a population. Fitdadi, like most methods used in population genetics, assumes the Wright-Fisher model of reproduction. It is currently unknown, however, whether using this model, which does not account for age structure, affects the inference of the DFE. Therefore, using SLiM, we simulated evolution over populations with various age structures. We then used the resulting SFSs to infer the DFEs using Fitdadi. We found that the inferred DFE was much more deleterious than the DFE used in our simulations. Our study shows that certain attributes of a population that are not present in Wright-Fisher models, such as age structure, may need to be accounted for when inferring the DFE using an SFS.
CHOTAI, STONE: Estimating migration rates in stepping-stone models
MEERA CHOTAI1, BEN STONE1, April Wei2,3,4, Sriram Sankararaman2,3,4
- BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
- Department of Computer Science, UCLA
- Department of Human Genetics, David Geffen School of Medicine, UCLA
- Department of Computational Medicine, David Geffen School of Medicine, UCLA
Demographic parameters such as migration rates and effective population sizes are key to understanding the genetic and evolutionary history of populations. Currently, there are no scalable methods that can accurately estimate asymmetric migration rates. We simulated population genetic data under a stepping-stone model using msPrime with symmetric and asymmetric migration. These migration rates were randomly sampled under a log10-uniform distribution between -4 and -2. We computed FST and site frequency spectrum from the simulated datasets as features and trained Lasso, Ridge, and random forest regression to predict the migration rates between adjacent subpopulations. We show that symmetric migration rates can be accurately predicted using any linear regression model (r2 = 0.99), and asymmetric migration rates are most accurately predicted with Ridge regression (r2 = 0.96). Our results suggest that population genetic simulations, combined with machine learning methods, can be powerful in unraveling complex demographic history and could be broadly applicable.
CHUNG: Investigating chromatin accessibility and transcriptomic changes in ASXL1-mutated fibroblasts
SAMANTHA CHUNG1, Leroy Bondhus2,3, Isabella Lin2,3,4, Valerie Arboleda2,3,4,5,6
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Human Genetics, UCLA
3 Department of Pathology and Laboratory Medicine, UCLA
4 Department of Computational Medicine, UCLA
5 Bioinformatics Interdepartmental Program, UCLA
6 Molecular Biology Institute, UCLA
Bohring-Opitz Syndrome (BOS) is a rare genetic syndrome caused by a heterozygous de novo mutation in ASXL1. We analyzed RNA and ATAC-seq data from fibroblast samples from BOS patients and controls. ATAC-seq identifies peaks with open chromatin accessibility and RNA-seq, identifies gene expression levels genome wide. We hypothesize that ASXL1 mutations drive changes in chromatin accessibility that alter specific gene expression. Using R, we mapped ATAC peaks to genes on the Human Gene GRCh38.13, and conducted statistical analyses and pathway enrichment analyses. 24 genes were found to be significant in both ATAC-seq analysis and RNA-seq (FDR < 0.05) . Gene ontology analysis for ATAC peaks (FDR < 0.05) showed both more open and closed chromatin near genes involved in morphogenesis and development. Gene ontology analysis for RNA-seq peaks shows functions associated with morphogenesis and development. These results may provide greater insight into understanding the pathophysiology of BOS. [video width="2560" height="1440" mp4="https://qcb.ucla.edu/wp-content/uploads/sites/14/2021/08/Chung.mp4"][/video] [/av_toggle] [av_toggle title='CHUNG, CRISANTO: Identifying tissue of origin using neural networks and RNA-seq data' tags='' custom_id='' av_uid='av-t8e314' sc_version='1.0'] JAY-HO CHUNG1, ALVARO CRISANTO1, Roy Wollman
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Integrative Biology and Physiology, Department of Chemistry and Biochemistry, UCLA
Neural networks offer a variety of uses as a method of data analysis. The simplest neural network can be used to identify digits from pictures from the Modified National Institute of Standards and Technology (MNIST) database. This project aimed to apply a neural network to biology by constructing a network which could identify tissue of origin from RNA-seq data. Data was downloaded from the GTEx Portal website and modified using the python programming language. The results indicate that the neural network did not identify the correct tissue of origin with a high-enough accuracy to warrant it as effective. However, the model has large potential such that with the correct modified data and model architecture, the model could increase its accuracy. Furthermore, the constructed model will serve as a template for future analysis of biological data where identification needs to occur.
COLE: Non-coding RNA regulation in breast cancer
NOLAN COLE1, Arjun Bhattacharya2, Bogdan Pasaniuc2
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Pathology & Laboratory Medicine, David Geffen School of Medicine, UCLA
Multiple expression quantitative trait loci (eQTLs) in protein coding genes (pc-genes) have been demonstrated to have a significant effect in the development of breast cancer – among other cancers. However, the overwhelming majority of these eQTLs locally regulate breast cancer-related genes. Distal effects of genetic variants on pc-genes have not been well-characterized. We investigate the distal effects that eQTLs have in non-coding RNA (ncRNA) and miRNA within solid-state breast carcinoma tumors. Using publicly available data from 437 European-ancestry patients in The Cancer Genome Atlas (TCGA), we optimize a statistical model for detecting cis/trans-eQTLs for both ncRNA and pc-genes. We will use these results in mediation analysis to quantify ncRNA influence on trans-eQTL effects. Preliminary results from a non-optimized model found 437 trans-eQTLs of protein-coding genes are cis-eQTLs of ncRNAs. After mediation analysis, we ultimately hope to find how ncRNAs or miRNAs affect pc-genes relevant in breast cancer development and progression.
DAUTLE: Predicting candidate genes for the development of therapies to treat focal amplifications in cancers
MADISON DAUTLE1, Kai Song2, Ashvath Balgovind3, Thomas Graeber3,4,5,6,7
- BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
- Department of Bioengineering, UCLA
- Department of Molecular and Medical Pharmacology, David Geffen School of Medicine, UCLA
- Jonsson Comprehensive Cancer Center, UCLA
- Crump Institute for Molecular Imaging, UCLA
- California NanoSystems Institute, UCLA
- Eli and Edythe Broad Center of Regenerative Medicine and Stem Cell Research, UCLA
Oncogenes amplified on extrachromosomal DNAs and homogenously staining regions – collectively referred to as focal amplifications – increase the heterogeneity of cellular genomes, decreasing treatment efficacy. This project seeks to identify candidate genes for experimental testing to treat focal amplifications in cancers. Using the R programming language, we performed gene expression and CRISPR screen analyses on the CCLE cell line, 21Q2 public dataset. A linear model dependent on focal amplification status, aneuploidy score, tissue type, and copy number variation was developed to reveal genes vital to focal amplification positive cells. The resulting list of ranked genes was investigated to identify candidate genes related to DNA repair, DNA replication, DNA tethering, vesicles, and genomic instability. These candidate genes will inform experimental research aimed at discovering and developing new therapies for the treatment of focal amplifications in cancers.
DIMOWO: Exercise alters bacterial composition in the gut microbiome from the Hybrid Mouse Diversity Panel
OIZA Y. DIMOWO1,2, Timothy M. Moore3, Arjen J. Cupido3,4, Aldons J. Lusis3,4,5
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Life and Physical Sciences, Fisk University, Nashville, TN, USA
3 Department of Medicine, Division of Cardiology, UCLA
4 Department of Vascular Medicine, Amsterdam UMC, University of Amsterdam, Netherlands
5 Department of Human Genetics, David Geffen School of Medicine, UCLA
6 Department of Microbiology, Immunology and Molecular Genetics, UCLA
The critical importance of the gut microbiome is becoming increasingly appreciated. Exercise is known to reverse and prevent complications associated with metabolic disorders. However, whether exercise is a potent modulator of gut microbiota composition and how this occurs is unknown. Here, we investigated the role of exercise on the bacterial composition of the gut in the Hybrid Mouse Diversity Panel. Shotgun sequencing and differential abundance testing of our samples revealed several taxa that were significantly altered in exercised animals. A better understanding of the effect of exercise on the gut microbiota would aid in the development of strategies to regulate microbial composition and diversity while ultimately enhancing human metabolic health.
EL OUSROUTI, TIMMONS: Assessing the impact of transcriptome annotations on isoform-level quantification in RNA-seq
HAMZA EL OUSROUTI1, SARA TIMMONS1, Minsoo Kim2,3, Michael Gandal2,3
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Psychiatry, David Geffen School of Medicine, UCLA
3 Department of Human Genetics, David Geffen School of Medicine, UCLA
Different choices of transcriptome annotations can have a substantial impact on downstream genomic analyses, such as variant effect prediction and gene expression quantification. Yet, it remains unclear how these annotations can affect isoform-level quantification in RNA-seq. Here, we systematically compare isoform-level expression in 120 human fetal brain samples across multiple transcriptome annotations. We find negligible discrepancy between isoform expression when quantifying expression based on different versions of GENCODE (release 33 vs 38), while we observe substantial discrepancy when filtering for transcripts with well-supported annotation. Due to the lack of the ground truth for isoform expression, it is hard to discern the more accurate quantification, but we expect such quantification to yield a larger eQTL discovery in the future. Overall, our findings highlight the need for more comprehensive and high-confidence transcriptome annotation for accurate isoform-level quantification.
EMEONYE: Assessing the preservation of somatic copy number alterations in glioma model systems
CHIDERA O. EMEONYE1, Nicholas A. Bayley2,3, Christopher Tse2, Henan Zhu2, David A. Nathanson2,4, and Thomas G. Graeber2,4.
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Molecular and Medical Pharmacology, UCLA
3 Bioinformatics Interdepartmental Program, UCLA
4 Jonsson Comprehensive Cancer Center, UCLA
Patient-derived model systems are increasingly used in translational cancer research as they are presumed to faithfully represent the genomic features of primary tumors. Recent studies have shown a divergence in the genomic and mutation profile of patient-derived models raising concerns that these models may not be fully representative of human tumors. Glioblastoma is the most common and aggressive primary brain tumor, often driven by somatic copy number events. Here we analyzed and compared copy number changes of glioblastoma patient tumors to their matched orthotopic xenograft or gliomasphere culture models to identify genomic sites and genes with the greatest divergence and found differing genomic evolution in the different model systems. Our findings suggest that different model environments impose different selective pressures on genomic alterations. Therefore, translational researchers need to consider their model system carefully and ensure genomic fidelity before testing targeted therapeutics in model systems.
ERCELEN: Assessing T-Cell Receptor Diversity and Genetic Variation in Complex Phenotypes Using Off-Target Sequencing
DEFNE ERCELEN1, Michael James Thompson, Noah Zaitlen
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
The diversity of the adaptive immune system is critical for maintaining and developing protective immunity. In particular, hypervariable complementary-determining regions (CDRs) in the genome determine the structure of immunoglobulins and T-cell receptors (TCRs). Nonetheless, the extent to which an individual’s immune repertoire is predetermined remains contested. We utilized IMREP, software designed to detect CDR regions, with off-target reads from whole exome sequencing (WES) data in the UKBiobank consortium. Confirming the validity of our approach, we observed relatively high concentrations of alpha and beta chains in comparison to delta or gamma chains. We also find significant correlations between the number and alpha diversity of TCRA and phenotypes such as living in an urban or rural setting. Finally, we conducted a GWAS on alpha diversity of TCRA receptors and found two statistically significant loci. Our results suggest immune signatures may not only distinguish between cases and controls, but that they may also be partly predetermined by genetics.
FISHER, JORDAN: Characterizing SNV Mutational Signatures in Cancer Samples
OLIVIA FISHER1,2,3,4,5, MADISON JORDAN1,2,3,4,5, Takafumi N. Yamaguchi1,2,3,4,5, Nicholas K. Wang1,2,3,4,5, Rupert Hugh-White1,2,3,4,5, Paul C. Boutros1,2,3,4,5
1 Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Human Genetics, University of California, Los Angeles
3 Department of Urology, University of California, Los Angeles
4 Institute for Precision Health, University of California, Los Angeles
5 Jonsson Comprehensive Cancer Center, University of California, Los Angeles
Mutational signatures within the genome are characteristic combinations of various mutations such as base substitutions, or indels. These signatures inform us of the process by which disruptive mutations lead to carcinogenesis. Single Nucleotide Variants (SNVs), the most common somatic mutation in the cancer genome, can promote tumor initiation, progression, and resistance to therapy. We aimed to assign SNVs to specific trinucleotide signatures, using their distinct patterns to quantify the level of mutational activity in each patient or tumor. We quantified signature activities in patients using high-performance computing to implement two existing algorithms within a Nextflow pipeline. The pipeline provides a cohesive framework to automate mutational signature detection and SNV assignment. Our results will characterize mutational signatures present in the genome to predict mutational activities in a sample and assign signature probabilities to specific SNVs. By labeling each patient’s genome with the most probable signature, tumors can be diagnosed at a molecular level and allow for predicting targeted treatment strategies.
FRASER: Identifying primary dysregulated genes in RelB-deficient dendritic cells
ANNA FRASER1, Héctor Navarro2, Diane Lefaudeux2, Yi Liu2, Alexander Hoffmann1,2
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Microbiology, Immunology, & Molecular Genetics, UCLAImmune response genes in innate immune cells are largely regulated by NFκB and IFN signaling.
However, the loss of one NFκB transcription factor subunit, RelB, has been shown to lead to detrimental inflammation and autoimmunity in both mice and humans. Preliminary data suggests this phenotype coincides with elevated Type I IFN signaling in dendritic cells (DCs). Here we aimed to identify genes that are dysregulated in RelB-deficient cells and that in turn might lead to hyper-active IFN signaling. We performed RNA-seq analysis on RelB-/- and Ifnar-/-RelB-/- DCs, as well as controls, stimulated with pathogen-derived molecules poly(I:C) and CpG in a 5-point timecourse. All samples passed rigorous quality control tests. Differential gene expression analysis revealed cohorts of genes, but surprisingly little evidence for interferon hyperactivity. Instead, the newly identified genes may suggest new research avenues to understand how RelB suppresses autoimmunity, leading to improved interventions for pathologies caused by RelB deficiency.
GARDNER: Using Bioinformatics Tools to Investigate Prevalence of Amino Acid Substitutions in Bacteria Expressing TEM Beta-lactamase Resistance Genes
SHEANEL GARDNER1,2, CYNTHIA WANG, Portia Mira2, Sada Boyd2, Pamela Yeh1
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Systems Biology, UCLA
Antibiotic resistance, a major threat to global health, occurs when bacteria evolve and mutate to confer resistance to antibiotics. Constant exposure to commonly used antibiotics like β-lactam antibiotics, has caused bacteria to produce an extended spectrum of beta-lactamase resistance genes, including TEM. Here, we investigate the prevalence of TEM genotypes in environments exposed to different β-lactam antibiotics and how this prevalence corresponds to bacterial growth. A 5-day competition experiment was conducted using equal proportions of all 16 possible mutations of the four amino acid substitutions found in TEM-85 variants exposed to different sublethal concentrations of beta-lactam antibiotics and serial transferred daily. Sanger sequencing was applied to just the TEM gene of the remaining bacterial populations at the end of the 5-day experiment. After calculating the frequency of each substitution and each variant genotype, we found that the growth rate trend of the genotypes was directly proportional to the genotype prevalence discovered. These results show that bacterial growth rates can be used to understand bacterial population dynamics in certain antibiotic exposed environments.
GLEASON: Fibroblast Added Guiding Phenotype Orchestrates Oral Barrier Tissue Immunity
ANNE GLEASON1, Takeru Kondo2, Ichiro Nishimura2
1BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2Weintraub Center for Reconstructive Biotechnology, UCLA School of Dentistry
Periodontitis is one of the most prevalent human inflammatory diseases; however, the underlying mechanism regulating disease progression is understudied. We used single-cell RNA sequencing to analyze gingival cells of mouse ligature-induced periodontitis at initiation (D1), progressive (D4), and fully developed (D7) stages. As expected, pro-inflammatory immune cells increased from D1 to D7, highlighted by a phenotype shift to Th17 cells and the accumulation of neutrophils. This study discovered a unique set of gingival fibroblasts, which strongly expressed toll-like receptors and chemokines. The newly found gingival fibroblasts with an added guiding phenotype of microbial-immune interaction were named ag-fibroblasts. Throughout periodontitis pathogenesis, ag-fibroblasts appeared to be responsible for sensing pathogens and recruiting immune cell responders through signaling interaction by chemokines and their putative receptors. The discovery of ag-fibroblasts presents a new avenue for future study to uncover the underlying disease mechanism and provides a novel clue in the development of potential therapeutics.
GRIFFIN: Influence of foreign genes via cell-cell fusion on Tumorigenesis
TAYLOR GRIFFIN1, Mengtao Li2, Yi-Ling Lin2
1BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Division of Diagnostic and Surgical Sciences, UCLA School of Dentistry
SCCVII/SF is a mouse squamous cell carcinoma line that forms tumors after injecting into syngeneic mice. Fusing this cell line with mouse bone marrow macrophages via the RANKL fusion agent was conducted to gain insight into the possible impact of this fusion on tumor growth. The SCCVII/SF-macrophage fusion cells were cloned and injected into mice. After two weeks, some clones produced no/small while others produced large tumors. The former clones were submitted for RNA-Seq and the data were analyzed via FastQC, trimmed with trimmomatic, and quality controlled. The in vivo tumor contents were also quantified using flow cytometry. Investigating differences between RNA-Seq data and cellular makeup of the control cell line and the SCCVII/SF-macrophage cell line that led to no tumor/small tumor phenotypes can help to better understand the functionality of differential gene expression in cancer genomics.
HE: Characterizing Temporal Patterns of Mutations in SARS-CoV-2 using ConsHMM conservation states
CHRISTOPHER HE1, Soo Bin Kwon2, Jason Ernst2
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Bioinformatics Interdepartmental Program, University of California, Los Angeles, CA, USA
New variants of SARS-CoV-2 are constantly being sequenced, providing a wealth of information on how the viral genome is changing over time. With the fast-evolving nature of the SARS-CoV-2 genome, the ability to predict the fitness of new strains is indispensable. Our lab recently used ConsHMM to generate an annotation of the SARS-CoV-2 genome based on multiple sequence alignment, providing novel information for interpreting the genome. This project analyzes mutational trends in the SARS-CoV-2 genome using the ConsHMM state annotations. We analyzed trends in mutational frequency over time based on ConsHMM state annotation and characterized ConsHMM states and genes based on enrichment of mutations. We found that state annotations were predictive of staying power of new mutations. Overall, we provided evidence that ConsHMM state annotations are useful in identifying mutational trends and thus may be able to contribute significantly to making predictions about future trends in SARS-CoV-2 evolution.
HERR: Evaluation of the genomic selection models on DNA Methylation
SAM HERR1 Matteo Pelligrini2
- BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
- Departments of Molecular, Cellular and Developmental Biology, UCLA
Phenotypic prediction has typically relied on genomic models that use SNPs to estimate trait values. While these models have had success, there remain limitations in their predictive ability. Here we ask whether combining genetic and epigenetic data can improve the accuracy of predictions generated by a library of established genomic models. We found that methylation models were not able to predict weight as well as genetic models but were able to detect locations of variable methylation levels that are strongly correlated with weight. However, the combined model of both methylation and genotype had a similar predictive ability as genotype alone. By contrast, we found that epigenetics alone has the most accurate prediction of age. Overall, we show that bayesian models work with continuous methylation data but combined models, at least for these two traits, do not improve predictions. Nonetheless, we expect traits with a strong genetic and epigenetic component may benefit from this combined approach.
HIRAKI: Genome-wide trans-eQTL scans in low-coverage RNA-Seq
HANNAH HIRAKI1, Tommer Schwarz2, Bogdan Pasaniuc3,4,5
1 B.I.G. Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Bioinformatics Interdepartmental Program, UCLA
3 Department of Pathology and Laboratory Medicine, UCLA
4 Department of Human Genetics, UCLA
5 Department of Computational Medicine, UCLA
Expression Quantitative Trait Loci (eQTLs) are polymorphisms that play a role in regulating gene expression of complex traits. They can be classified as either cis (proximal) or trans (distal) depending on the location of the associated gene with respect to the associated variant. Previous studies have identified associations between disease-related expression and cis-eQTLs, but few have produced substantial numbers of trans-eQTLs due to small effect sizes and limited sample sizes. For this project we utilize low-coverage RNA-Seq from whole blood to optimize a pipeline for the discovery of trans-eQTLs. By investigating the functional enrichment of the eGenes identified from trans-eQTL analysis, we hope to gain insight into their mechanisms of regulating disease pathways.
HUGGINS: Using Variation Analysis for the Identification of Mutations Linked with Antibiotic Resistance in Staphylococcus Epidermidis
CASEY HUGGINS1, Natalie Lozano-Huntelman2, Portia Mira2, Pamela Yeh2
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Ecology and Evolutionary Biology, UCLA
Living cells have continually adapted to changing environments since the dawn of life and moving into the intercellular warfare that brought about chemical antibiotics. In combating bacterial infections, antibiotics such as piperacillin and tetracycline are typically used to eliminate the populations of bacteria wreaking havoc. However, bacterial cells have adapted numerous ways to combat the variety of chemical agents that inhibit their cellular processes. Provided a limited variety of antibiotic options, it is more important than ever to understand the processes by which these resistances come about. In this study, strains of Staphylococcus epidermidis were selected for resistance to piperacillin and tetracycline. Using whole genome sequencing and variation analysis, conserved mutations were found for each drug resistance. By identifying the mutations responsible for resistance to particular antibiotics, networks of cross resistance and even collateral sensitivity could be constructed which may then allow for better courses of antibiotics in clinical applications.
IKECHUKWU, SEHAJMEET: A Deep Learning Model for Segmenting Nuclei on Digital Histopathology Images
MOSHE IKECHUKWU2, SEHAJMEET SOHAL2, Dr. William Hsu1
1 BIG Summer Program, Department of Radiological Sciences, UCLA
2 BIG Summer Program scholar 2021
Digital pathology is the acquisition of high-resolution histology slides that can be analyzed using computational methods. Issues with staining and scanning across the sites and biological variance in the slides make the task of segmenting nuclei is extremely challenging. Our goal is to create a model using deep learning that differentiates between non-small cell lung cancer subtypes. U-Nets have been shown to accurately segment images. Hence we used a U-Net model for nuclei segmentation. The model was trained and tested using data from the 2018 Kaggle Data Science Bowl (670 pairs [images+masks] for training and 65 pairs for testing) and data from Janowczyk et al (141 pairs for training and 28 pairs for testing). To evaluate our model, Dice coefficient and intersection over union (IOU) were used to compare our segmentation results with the human-generated reference. Initial results demonstrate that U-Nets are capable of performing this task. We will eventually use the model to quantify nuclei characteristics towards differentiating adenocarcinoma and squamous cell carcinoma.
JAFFE: Differential gene expression and weighted gene correlation network analysis in a heart failure with preserved ejection fraction mouse model
MAYA Y. JAFFE1, Arjen J. Cupido2,3, Timothy M. Moore4, Aldons J. Lusis2,4,5
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Dept. of Medicine, Division of Cardiology, UCLA, Los Angeles, CA, USA
3 Dept. of Vascular Medicine, Amsterdam University Medical Centers, University of Amsterdam, Amsterdam, Netherlands
4 Dept. of Human Genetics, UCLA, Los Angeles, CA, USA.
5 Dept. of Microbiology, Immunology and Molecular Genetics, UCLA, Los Angeles, CA, USA
Heart failure (HF) prevalence in the US continues to rise, with approximately half of all HF patients having heart failure with preserved ejection fraction (HFpEF). The incidence of HFpEF is expected to increase, partly due to prevalent comorbidities such as obesity, diabetes, and hypertension. However, few effective therapeutic strategies for HFpEF have been identified. To facilitate the identification of therapeutic targets, we aimed to understand changes in gene expression in cardiac tissue from a HFpEF mouse model. We utilized RNA sequencing data from a HFpEF mice and control mice, and performed differential gene expression (DESeq package, R) and weighted gene correlation analysis (WGCNA package, R) and integrated the results. We observed that many of the most highly significant differentially expressed genes relate to fatty acid oxidation. When comparing significant genes within important modules, our results indicate that catabolism, muscle development, and angiogenesis play an important role in the HFpEF phenotype.
JAMAL: Integrated Reference and Cell Type Classification Pipeline Using Machine Learning Models
ZOEB JAMAL1, Justin Yee, Russell Littman2, Xia Yang3
1: BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2: Bioinformatics Interdepartmental PhD Program, UCLA
3: Interdepartmental Program of Bioinformatics, UCLA
Single-Cell RNA Sequencing enables profiling of individual cell transcriptomes, allowing assessment of transcriptional landscapes of cell populations However, identifying marker genes and labelling cell types can be laborious. We created an integrated reference dataset that can be used to train machine learning models to classify cell types and evaluated their performance. We integrated the Mouse Cell Atlas, Tabula Muris, and the Allen Mouse Brain Atlas and optimized hyperparameters for Support-Vector Machines and Naive Bayes models. Our results indicate that including more principal components during dimensionality reduction, tailoring hyperparameters to the reference, and using tissue-specific classifiers increases prediction accuracy. To benchmark, we calculated accuracy and compared clustering patterns of predicted cell types with those derived from manual annotation. We aim to build a web tool that allows users to upload data and receive classifications, which is an essential step in the study of physiology and disease.
KATZ, QIN: MiRNA-seq Analysis for the Study of Craniofacial Bone Regeneration
TYLER KATZ1#, JUDY QIN1#, Jiabing Fan2, Min Lee2
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Division of Advanced Prosthodontics, School of Dentistry, UCLA
# Authors contributed equally
The use of exosomes presents a promising cell-free approach for promoting bone regeneration. We previously reported that the implementation of exosome mimics (EMs) derived from genetically-modified mesenchymal stem cells (MSCs) exhibited significant craniofacial bone regeneration and growing evidence shows that miRNAs play an important role throughout the process. To elucidate the underlying mechanisms of EM-mediated osteogenesis, we investigated the profile of miRNA expression within EMs via miRNA sequencing. The data identified bundles of miRNAs that were up-regulated or down-regulated in different groups of EMs, which were then visualized using volcano plots. Both the Venn diagram and heatmap created further revealed that several miRNAs commonly expressed in pairwise comparison exerted synergistic up-regulation or down-regulation. MiRNA prediction analysis also indicated that the identified specific miRNA may interact with the BMP signaling pathway. Our work offers the foundation to develop further mechanism studies and ultimately, an exosome-based therapeutic approach for craniofacial bone regeneration.
KOELLING, LIU, PHAM: Modeling gene-gene interactions in GWAS data using deep learning and machine learning approaches
HENRY KOELLING1, NICHOLAS LIU1, TRONG PHAM1, Boyang Fu2, Sriram Sankararaman2,3,4,5
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Computer Science, UCLA
3 Department of Human Genetics, David Geffen School of Medicine, UCLA
4 Department of Computational Medicine, UCLA
5 Bioinformatics Interdepartmental Graduate Program, UCLA
Estimating the heritability of traits has become an important area of research in computational genetics – however, it remains a challenge to model gene-gene interactions (epistasis), which are suspected to contribute to the “missing heritability problem”. One current approach to detect such interactions employs a neural network to calculate Shapleyinteraction values between hidden nodes, which represent gene-level interactions. In this work, we evaluated the power and statistical calibration of this NN model and identified constraints to the model based on noise, sample size, and other hyperparameters. For datasets with limited samples, we propose a gradient-boosting decision-tree model, which we found to be better calibrated when detecting SNP-level interactions. By investigating the limitations of existing approaches to modeling epistasis and developing complementary, alternative models, we can better understand the source of missing heritability and the genetic architectures underlying complex traits, leading to developments in phenotype prediction and precision medicine.
LANDGREN, WENGLER: Quantifying the Read Depth Sensitivity of RNA-Seq Analyses in Prostate Cancer
ELISABETH LANDGREN1,2,3,4,5, JAMES WENGLER1,2,3,4,5, Ruilian Zhang 1,2,3,4,5, Samuel Shenoi1,2,3,4,5, John Lee1,2,3,4,5, Julie Livingstone1,2,3,4,5, and Paul Boutros1,2,3,4,5
1 Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Human Genetics, University of California, Los Angeles
3 Department of Urology, University of California, Los Angeles
4 Institute for Precision Health, University of California, Los Angeles
5 Jonsson Comprehensive Cancer Center, University of California, Los Angeles
RNA sequencing (RNA-Seq) is an established way of analyzing the transcriptome of cancer cells. RNA-Seq can inform us about transcript type and quantity, as well as, the abundance of fusion and alternative splicing isoforms, which provides insight into the behavior of cancer cells. An open question is how downstream results change based on sequencing depth. To address this question, we used RNA-Seq from 20 localized prostate tumors that were sequenced using paired-end sequencing to an average depth of 180 million reads. We down sampled each sample to seven different depths ranging from 5-120 million reads with multiple technical replicates. In total, we produced 680 samples that were processed through systematic pipelines. Understanding the differences in information in relation to sequencing depth can help researchers choose a sequencing depth to avoid false negatives and prioritize expenditures.
LIU Z: Benchmark of different isoform quantification methods in a simulation-based framework
ZHENGTON LIU2, Wenbin Guo1, Xinzhou Ge3, Jingyi Jessica Li3
1 Bioinformatics IDP, UCLA
2 BIG Summer Program, Institute of Quantitative and Computational Biosciences, UCLA
3 Department of Statistics, UCLA
Methodological advances have greatly improved the accuracy of isoform quantification from short-read RNA-seq data. With different isoform quantification tools proposed, multiple studies have leveraged the isoform abundances to study the isoform abundance quantitative trait loci (isoformQTL). However, these studies exhibit a great disparity in their choice of methods for isoform quantification and association testing, and there have not been any benchmark on these isoformQTL studies. Therefore, this project aims to generate simulated datasets and compare existing isoformQTL methods by applying them to our simulated dataset. We use the isoform abundance data generated from the GTEx real data to guide our simulation design. As we have the truth for simulated datasets, we can benchmark isoformQTL methods in terms of their precision and power for identifying true isoformQTLs. Our current results indicate that Kallisto, Salmon and Cufflinks achieve better performance than RSEM on both the simulated and GTEx real data sets.
LY, WEBER: Non-invasive Identification of Bacterial exRNAs for Early Detection of Gastric Cancer in Human Saliva
CARA LY1, RYAN WEBER1, Karolina Kaczor-Urbanowicz2, David T. Wong2
- BIG Summer Program, Institute of Quantitative and Computational Biosciences, UCLA
- UCLA School of Dentistry, Center for Oral/Head & Neck Oncology Research, UCLA
Gastric cancer (GC) is the fourth most common cancer diagnosed globally and the third leading cause of cancer-related deaths due to lack of early symptoms. Previously, bacteria have been associated with the development of GC. As saliva is a non-invasive biofluid containing protein, RNA, and microbial communities, we aim to identify bacterial exRNAs associated with gastric cancer in human saliva. We analyzed the salivary short RNA sequencing profile of 10 GC samples and 10 controls using the exceRpt pipeline and DeSeq2 in order to investigate the microbial communities. Our preliminary data revealed 18 statistically significantly differentially expressed bacterial species between 5 gastric cancer and 5 control samples (padj < 0.05) including Prevotella nigrescens (padj=2.1E-07) and Pediculus humanus (padj=2.3E-03). Identification of salivary biomarkers for GC would allow for the development of a non-invasive and cost effective screening tool that could lead to earlier detection of GC. [video width="2560" height="1440" mp4="https://qcb.ucla.edu/wp-content/uploads/sites/14/2021/08/Ly-Weber-1.mp4"][/video] [/av_toggle] [av_toggle title='MANZANO: Using Cell-Free DNA Methylation Sequencing Data for Cancer Detection' tags='' custom_id='' av_uid='av-30hgk8' sc_version='1.0'] JOMEL MANZANO1, Wenyuan Li2, Xianghong Zhou2
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Pathology and Laboratory Medicine, David Geffen School of Medicine, UCLA
The detection of cell-free DNA (cfDNA) in plasma has led to major applications in cancer detection. However, identifying the tiny amount of circulating-tumor cfDNA poses a major computational challenge to cancer diagnosis. This study applies the CancerDetector algorithm to address this challenge by introducing a statistical model that can increase the signal-to-noise ratio. The model uses the cfDNA methylation sequencing data, classifies individual cfDNA reads to be either from tumor cells or normal cells and uses these categorized reads to estimate the tumor burden in plasma. We implemented this algorithm in R, ran this model on simulated data, and calculated a precise tumor burden given the cfDNA methylation data and cancer markers. This method is generalized to diagnose other diseases and is implemented in this study. This R implementation can benefit the research community of cfDNA research and be applied to the data of real patients.
MANZANO: Using Cell-Free DNA Methylation Sequencing Data for Cancer Detection
JOMEL MANZANO1, Wenyuan Li2, Xianghong Zhou2
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Pathology and Laboratory Medicine, David Geffen School of Medicine, UCLA
The detection of cell-free DNA (cfDNA) in plasma has led to major applications in cancer detection. However, identifying the tiny amount of circulating-tumor cfDNA poses a major computational challenge to cancer diagnosis. This study applies the CancerDetector algorithm to address this challenge by introducing a statistical model that can increase the signal-to-noise ratio. The model uses the cfDNA methylation sequencing data, classifies individual cfDNA reads to be either from tumor cells or normal cells and uses these categorized reads to estimate the tumor burden in plasma. We implemented this algorithm in R, ran this model on simulated data, and calculated a precise tumor burden given the cfDNA methylation data and cancer markers. This method is generalized to diagnose other diseases and is implemented in this study. This R implementation can benefit the research community of cfDNA research and be applied to the data of real patients.
MCQUILLEN: Single-Nuclei Analysis of Sex Differences in Mouse Medial Septum
CADEN N. MCQUILLEN1, Montgomery Blencowe2, Graciel Diamante2, In-Sook Ahn2, Jason Lerch5, Allan Mackenzie-Graham4, Armin Raznahan6, Art Arnold2, Xia Yang2,3
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Integrative Biology and Physiology, UCLA
3 Interdepartmental Program of Bioinformatics, UCLA
4 Department of Neurology, UCLA
5 Nuffield Department of Clinical Neuroscience, The University of Oxford
6 Developmental Neurogenomics Unit, Human Genetics Branch, NIMH
Humans display robust sex differences in neurodevelopment starting from early childhood through to adulthood. Consequently, these differences result in sex-specific biases towards many neurological disorders including autism, ADHD, and major depression disorder. The mechanisms driving these biases are largely believed to be caused by two biological differences: the effect of gonadal hormones (gonadal effect) and the presence/absence of the Y chromosome (sex chromosome effect) but uncovering their unique contribution in human health and disease is challenging. Therefore, our study aims to analyze these sex driven effects at a single-nuclei resolution using the 8-core genotype model, which not only can distinguish the difference between the gonadal and chromosomal effect but also provides critical insight to a chromosomal dosage effect. With this we compared the transcriptome of 8 genotypes of mice (XXF, XXM, XYF, XYM, XXYF, XXYM, XYYF, XYYM) highlighting changes across cell populations, gene expression, and the relevance of these changes to disease by enrichment analysis for over 70 disease traits. Notably we observed many psychiatric related disease traits to be strongly enriched in myelinating oligodendrocyte populations, which was further magnified when examining the chromosomal dosage effect suggesting the importance of both the composition and number of sex chromosomes in sex biased neurological disorders.
MELGOZA: Simulating Effects of a T-Type Ca2+ Current in Realistic Neuron Models
ITZEL MELGOZA1, Sharmila Venugopal2
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Integrative Biology and Physiology, UCLA
Neurons express diverse voltage-activated Ca2+ channels on their plasma membrane which contribute to their membrane potential, spiking and circuit functions. Transient or T-type Ca2+ channels are low-voltage activated and permit sustained Ca2+ influx near a neuron’s resting membrane potential. Different isoforms of the T-type channel Ca2+ currents expressed in a thalamocortical circuit are posited to underlie the spike-and-wave discharges in absence epilepsy. As these isoforms are difficult to target pharmacologically, a computational model of the channel could assist in studying the channel biophysics and the resulting physiological behavior of thalamocortical neurons. Using the NEURON software, we show that increasing the conductance of a T-Type Ca2+ current transforms tonic spiking to burst discharge in a model neuron. As a cortical neuron displayed physiological behavior characteristic of T-Type channels, this model can be used to study the role of T-Type current isoforms during epileptic discharges in real and model neurons.
MOORTHY, YOO: Exploring Cell-Cell Interactions with Hypergraph Representation Learning
AHILA MOORTHY1, HYERY YOO1, Yihe Deng2, Quanquan Gu2
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Computer Science, UCLA
Cell-cell interactions (CCIs) are vital to the functioning and development of multicellular organisms. With recent development of single-cell RNA sequencing techniques, CCIs can be detected based on the co-expression of ligand-receptor (L-R) pairs. Such interactions could be commonly represented with graphs, where various analysis methods are developed for the data structure. However, as the interactions between cells are diverse and go well beyond binary, it could be more efficiently represented as a hypergraph. In this work, we are interested in exploring CCIs from this less studied perspective. We propose a hypergraph representation learning framework based on Hyper-SAGNN to capture and study the complex interactions between cells. We demonstrate empirically that our method could accurately predict the interactions and generate useful representations for further analysis. Potential applications of this research include a better microscopic understanding of the functions of the immune system, tissue organization, and cell growth-based diseases such as cancer.
MOSQUERA, SHODUBI: Investigating the effect of immunosuppressive drugs on macrophages through RNA-seq
SAMUEL MOSQUERA1, OLUWAPELUMI SHODUBI1, Quen Cheng2
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Medicine, Infectious Diseases, UCLA
Transplant patients are treated with immunosuppressive drugs to prevent transplant rejection by the adaptive immune system. However, the commonly used steroid hormone prednisone and the non-steroid tacrolimus also inhibit the function of innate immune cells, increasing the risk for fungal infections. Here, we analyzed innate immune macrophage transcriptomes upon exposure to the fungus Aspergillus or the fungal molecule beta-glucan, in the presence or absence of Tacrolimus or Prednisone. We evaluated the quality of RNA-sequencing datasets generated at various time points. Focusing on datasets that passed rigorous quality control metrics, our results suggested that tacrolimus did not significantly affect macrophages while prednisone directly diminished the expression of cytokine genes. These results suggest that prednisone but not tacrolimus impairs macrophage function to combat fungal infections in transplant patients, motivating a refinement of current immunosuppression protocols.
NGUYEN: Detecting Differentially Expressed Genes along Single-cell Pseudotime through QGAM
HUY NGUYEN1,2, Tianyang Liu2, Melody Zhang2
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 The Junction of Statistics and Biology, UCLA
To understand molecular mechanisms behind cell state changes, one crucial step is identifying differentially expressed (DE) genes along inferred single-cell pseudotime. Currently, many existing methods do not provide accurate detection when the gene counts data contains many outliers. In this study, we simulated single cell sequencing data with low, medium, and bifurcation dispersion. Next, we used the advanced fast calibrated Bayesian methods for fitting quantile GAMs (QGAMs) to demonstrate the effectiveness of the method in distinguishing DE genes and non-DE genes. More so, we benchmarked the method with two other state-of-the-art methods- PseudotimeDE and tradeSeq- to underline the robustness of QGAM. The results showed that QGAM had the lowest False Discovery Rate, most uniform distribution of non-DE genes’ p-values, and good AUC scores. As QGAM is a relatively fast, new, and versatile technique, the study’s results can truly promote its uses in DE genes detection and beyond.
OKENWA, PETERSON: Introducing a Sparse k-Means Clustering Algorithm for Analyzing Genetic Data
CHIDERA OKENWA1, DANIEL PETERSON1, Benjamin Chu2, Seyoon Ko4, Hua Zhou4, Kenneth Lange2,3,4,5
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Biomathematics, David Geffen School of Medicine, UCLA
3 Department of Human Genetics, David Geffen School of Medicine, UCLA
4 Department of Biostatistics, David Geffen School of Medicine, UCLA
5 Bioinformatics Interdepartmental PhD Program, UCLA
Ancestry is a confounding factor in genome-wide association studies (GWAS). There are many algorithms to explore genetic data. The sheer quantity of data creates a need to develop dimension-reducing algorithms to analyze this data. We present a simple, yet effective k-means clustering algorithm (SKFR) that clusters genetic data based on most informative genetic markers (SNPs) among samples. This algorithm maintains low computational complexity while delivering a sparse solution. Using Julia programming, we analyzed the effectiveness of SKFR by comparing its results to results from other ancestry-determining methods as well as to true ancestry origins of samples. We found that through the SNP-ranking of SKFR, it accurately clusters people into their subpopulations even with a relatively low number of SNPs included. These results reveal SKFR is a computationally simple and effective unsupervised learning algorithm that can quickly process genetic data to deliver more accurate ancestry results and aid future genetic research.
ORELLANA, PIMENTEL-SOLORIO: Exploring the Role of Genetic Ancestry on Severity of Disease in Patients with Valley Fever
DIEGO ORELLANA1, GIOVANI PIMENTEL-SOLORIO1, Sarah Spendlove2,4,5,6, Manish Butte3, Valerie Arboleda2,4,5,6
- BIG SUMMER PROGRAM, David Geffen School of Medicine, UCLA
- Interdepartmental Bioinformatics Program, David Geffen School of Medicine, UCLA
- Division of Allergy and Immunology, Department of Pediatrics, David Geffen School of Medicine, UCLA
- Department of Pathology and Laboratory Medicine, David Geffen School of Medicine, UCLA
- Department of Human Genetics, David Geffen School of Medicine, UCLA
- Department of Computational Medicine, David Geffen School of Medicine, UCLA
Coccidioidomycosis, known as “Valley Fever”, is a fungal infection endemic to the Southwestern US. While most individuals have uncomplicated valley fever (UVF), a subset of patients progress to more severe disease, disseminated coccidioidomycosis (DCM). Our goal is to identify any genetic components that increase DCM risk compared to individuals with uncomplicated valley fever (UVF). We analyzed exome and genome data of Valley Fever patients obtained from UC Davis and the Valley Fever Institute using Unix, R, and Python. We performed quality control to match expected and observed sex and ancestry. In total, we have data from 561 individuals (DCM=159, UVF=342, Other=60). Our genetic data confirmed that individuals who self-identified as non-white had increased number of singleton SNPs by our quality control. We are performing additional analysis between phenotypes to determine whether they are associated with disease severity. Our findings will allow us to better understand the relationship between ancestry-specific gene regulation of the immune system and vulnerability to DCM.
PAYAN-MEDINA, WATSON: Assembly of an interactive database application for tracing atomic fates in metabolic networks
ADRIANA PAYAN-MEDINA1, CALEB WATSON1, Edward Ma2, Richard Law2, and Jun Park2
1BIG Summer Program, Institute of Quantitative and Computational Biosciences, UCLA
2Department of Chemical and Biomolecular Engineering, UCLA
Isotope tracers are indispensable for elucidating metabolic fluxes for the understanding and manipulation of cellular function. Tracing atoms across reactions is critical towards the design and interpretation of informative isotope tracing experiments. However, complex metabolic networks make it difficult to anticipate the fate of labeled atoms. Here, we introduce the interPathway Atomic Tracing Hub (iPATH), an application that interactively displays isotope tracing across a dynamic list of cellular systems and pathways. iPATH extends existing models by integrating resources into a consolidated database. Reaction atom transitions from this database are reindexed by pairwise comparisons of molecule topology, allowing atoms to be accurately traced across a network. From diagnostic mapping of basic biochemical pathways in E. coli central carbon metabolism, iPATH aims to accommodate advanced biological networks across widely divergent organisms with faster query processing than other methods. This applicable framework makes iPATH a practical atom mapping tool to optimize tracing experiments.
PORTA: Identifying Cell Type Interaction eQTL Effects using Low-Coverage Bulk RNA-seq Data
ROSE PORTA1, Hannah Hiraki1, Nolan Cole1, Tommer Schwarz2, Bogdan Pasaniuc2,3,4,5
1 B.I.G. Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Bioinformatics Interdepartmental Program, UCLA
3 Department of Pathology and Laboratory Medicine, UCLA
4 Department of Human Genetics, UCLA
5 Department of Computational Medicine, UCLA
Expression Quantitative Trait Loci (eQTLs) are genetic variants that influence gene expression levels. eQTL analysis at the cell type-specific level can help to pinpoint the specific biological pathways leading to phenotypic variation for complex traits. However, previous studies on cell type-specific eQTL effects have leveraged single-cell RNA-seq or bulk RNA-seq on purified cell types, which is resource intensive. In this project, we used the software Cibersort to estimate cell type proportions from whole blood bulk RNA-seq data and Decon-eQTL to identify cell type interaction eQTL effects. We identified 253 significant cell type interactions across 20 cell types. Additionally, we found cell type interactions enriched in different pathways. Developing a more nuanced understanding of eQTL effects on a cell type-specific level will help to identify the roles of these variants in complex disease pathways, which will lead to more targeted and effective treatments.
POWELL: Integrating Data-Driven and Mechanistic Approaches in Cell Regulatory Pathway Analysis
ASHLYN POWELL1, Farnaz Mohammadi2, Aaron Meyer2
1 BIG Summer Program, Institute of Quantitative and Computational Biosciences, UCLA
2 Department of Bioengineering, UCLA
Though effective therapies exist for melanoma, resistance to these drugs inevitably develops. Previous studies have shown that resistance arises from rare cancer cells that are reprogrammed from a pre-resistant state. Several genes, including EGFR, NGFR, and AXL, are disproportionately expressed in pre-resistant cells and have been comprehensively profiled through knockout models and gene expression measurement. However, the broader regulatory events by which a cell enters this rare state are unclear. A unified model for how these components interact would help uncover drivers of this process. We built an ordinary differential equation model of the concentrations of mRNA corresponding to pre-resistant genes. We used this as a data-driven framework to identify gene-gene interactions by allowing all possible interactions, then comparing to gene expression measurements from each knockout using optimization implemented in Julia. The interaction parameters inferred by the model can be used to identify key regulators driving melanoma drug resistance development.
PUTNEY: Utilization of regression analysis to uncover pathways relevant to maximizing ferroptosis sensitivity in uveal melanoma
NICHOLAS PUTNEY1, Kim Paraiso2, Arpi Beshlikyan2, Thomas Graeber2,3,4,5,6,
1 BIG SUMMER Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Molecular and Medical Pharmacology, UCLA, Los Angeles, CA
3 Jonsson Comprehensive Cancer Center, UCLA
4 Crump Institute for Molecular Imaging, UCLA
5 California NanoSystems Institute, UCLA
6 Eli and Edythe Broad Center of Regenerative Medicine and Stem Cell Research, UCLA
Uveal melanoma (UVM) is the most common form of adult ocular malignancy, with about 7000 individuals being diagnosed worldwide each year. Within the newly diagnosed cases, roughly half are effectively treated by current therapies, while the other half eventually progress to a metastatic state. While there are currently no FDA-approved treatments for metastatic UVM, our lab has recently found a subset of UVM that is sensitive to ferroptosis induction via pro-ferroptotic drugs such as RSL3, a molecule that induces ferroptosis through the inhibition of glutathione peroxidase. Although pro-ferroptotic drugs are a promising treatment for metastatic UVM, there are varying degrees of ferroptosis sensitivity across instances of UVM. To address the problem of varying sensitivity, we conducted a linear regression analysis to uncover genes relevant to maximizing ferroptosis sensitivity in uveal melanoma. Targeting pathways that increase pro-ferroptotic sensitivity could increase the effectiveness of drugs that target this pathway, such as RSL3, leading to improved patient outcomes in metastatic UVM.
RAMIREZ: Projective Nonnegative Matrix Factorization (PNMF) incorporated into Surrogate Variable Analysis (SVA) for an Interpretable Surrogate Variable Analysis
LUCIA RAMIREZ1, Kexin Li2, Jingyi Jessica Li2,3,4
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Statistics, UCLA
3 Department of Biomathematics, David Geffen School of Medicine, UCLA
4 Department of Human Genetics, David Geffen School of Medicine, UCLA
Surrogate Variable Analysis (SVA) provides insight into the effects of unmodeled or hidden factors on gene expression data. There are a variety of SVA algorithms, including the Two-Step SVA, the Iteratively Reweighted SVA, and the Iteratively Adjusted SVA (IA-SVA) that consider different approaches, but these methods don’t provide much interpretation of the surrogate variables estimated. The aim of the project was to incorporate the Projective Nonnegative Matrix Factorization (PNMF) algorithm into SVA to provide biological interpretation. Single-cell RNA-Sequencing (scRNA-seq) data was simulated from published datasets to validate the results of an interpretable SVA algorithm since the true simulated hidden factors are known. Published scRNA-seq data was used to test the algorithm on, compare the performance to other variations of the SVA algorithm, and provide a basis for the simulated data. The results provided insight into which gene groups were enriched in each estimated hidden factor through GO enrichment analysis, providing the interpretation of the surrogate variables.
RAZAVILAR, ZHOU: Investigating the Origin of Unmapped RNA-Sequencing Reads in Whole-Blood Samples from Patients with Bipolar Disorder or Schizophrenia
AIDA RAZAVILAR1, *, JENNIFER ZHOU 2,*, Toni Boltz 3, Tommer Schwarz 4, Merel Bot 5, Roel Ophoff 3,5
1 Department of Biological Sciences – Neuroscience, Columbia University, NYC, NY, USA
2 Department of Biomedical Engineering, UT Austin, Austin, TX, USA
3 Department of Human Genetics, University of California, Los Angeles, CA, USA
4 Department of Bioinformatics, University of California, Los Angeles, CA, USA
5 Center for Neurobehavioral Genetics, University of California, Los Angeles, CA, USA
* These authors contributed equally to this work
The microbiota-gut-brain-axis is of interest for its demonstrated role in neuropsychiatric disorders. Contrary to the understanding of the blood as a sterile environment there is in fact the presence of an endogenous microbiome. We investigated the origin of RNA sequencing reads that remained unmapped to the human genome following STAR alignment and expected differences in the distribution between affected individuals and controls. RNA reads were extracted from 1944 individuals with schizophrenia (n=229), bipolar (n=1129), families of bipolar individuals, and healthy controls. Using the Read Origin Protocol, sequences were cross compared to microbial, fungi, immune, protozoa, and viral genomic material across groups. Results revealed lower levels of immune RNA and viral RNA reads in individuals with Schizophrenia, which was not observed in the Bipolar cohort. This continues to support the role of the microbiome and immunity in Schizophrenia specifically, and points towards a simple, potentially translational predictive measure within neuropsychiatry.
SMALL: The Practical Implementation of the Sensitive Cell-Free DNA Mutation Calling Method cfSNV
COLIN SMALL1, Shuo Li2, Wenyuan Li2, Xianghong Zhou2
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Pathology and Laboratory Medicine, David Geffen School of Medicine, UCLA
cfSNV is a software package providing hierarchical mutation calling and tumor cluster mutation profiling for tumor-derived cell-free DNA. The software offers state of the art performance in mutation calling especially for early-stage cancer patients who may still have low tumor burdens, but practical use of the software is currently hindered by long runtimes for whole-patient-scale data. By implementing several essential modules of cfSNV in C++, we achieve runtime reductions ranging from 35.3% to 94.9% as compared to cfSNV’s original Python implementation. Runtime reductions depended on the contents and operations of each module, with modules that perform many algebraic and statistical operations in loops benefiting more from translation than those performing simple sequential steps. However, cfSNV calls these modules many times per inference, scaling even modest improvements in single-run runtime to substantial savings in aggregate.
SPARKS: Decoding the Regulatory Landscape of LncRNAs in Cellular Senescence
WILLIAM M. SPARKS1, Mithun Mitra1,3, Huiling Huang2, and Hilary A. Coller1,3
1Department of Molecular, Cell, and Developmental Biology, UCLA
2Bioinformatics Interdepartmental Program, UCLA
3Department of Biological Chemistry, David Geffen School of Medicine, UCLA
Cellular senescence is a state of irreversible cell cycle exit and is associated with aging. Here we focused on the relatively unexplored regulatory activities of long non-coding RNAs (lncRNAs) in senescence. Based on RNA-seq analysis, 67 lncRNAs changed significantly in expression in both replication stress-induced and oncogene activation-induced senescence models of human lung fibroblasts. The majority of these lncRNAs (54/67 or ~81%) do not change with quiescence, a state of reversible cell cycle arrest, implying cell state specificity. Seventeen of these senescent lncRNAs were predicted to participate as competing endogenous RNAs by “sponging” miRNAs, and potentially leading to the upregulation of miRNA target genes involved in secretory pathways. Further analysis showed that 20 out of 67 lncRNAs may also regulate genes by forming RNA-DNA triplexes in specific gene promoter regions and acting as a scaffold for transcription factor binding. Taken together, our results indicate multiple regulatory pathways involving lncRNAs in cellular senescence.
UMUNNA: Rewiring of host metabolism by lytic vs temperate viral communities
Rewiring of host metabolism by lytic vs temperate viral communities
CHIBUNDU UMUNNA1, Nickie Yang1, Benjamin Knowles2
1BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2Department of Ecology and Evolutionary Biology, UCLA
Viral infections could be lytic or temperate depending on the host’s environment and physiological state. Viruses commonly rewire host cell metabolism upon infection, but it is unknown how that rewiring changes between lytic vs temperate infections. Here we analyzed viral metagenomes from healthy to degraded coral reefs across a spectrum of lytic to temperate viral communities to determine what functions were over- or under-represented in temperate viruses, focused on carbohydrate metabolic genes. Carbohydrate genes increased from lytic to temperate viral communities. At the pathway level for carbohydrate catabolism, Entner-Doudoroff pathway genes were over-represented in temperate viruses, while pentose phosphate pathway and glycolysis genes were more often under-represented. These results help us understand how viruses are rewiring cellular carbon metabolism under different conditions and highlights the need to holistically consider metabolic and physiological impacts of viruses within ecological systems.
VARMA: A robust measure of tissue specificity of gene expression
ROSHNI VARMA1,2, Yenifer Hernandez3, Leroy Bondhus4,5,6, Valerie Arboleda4,5,6
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Microbiology, Immunology, and Molecular Genetics, UCLA
3 Computational and Systems Biology Interdepartmental Program, UCLA
4 Department of Human Genetics, David Geffen School of Medicine, UCLA
5 Department of Pathology and Laboratory Medicine, David Geffen School of Medicine, UCLA
6 Department of Computational Medicine, David Geffen School of Medicine, UCLA
Tissue specificity is the degree to which a gene’s expression is restricted to a limited set of tissues. Measures of specificity can help explain the variable phenotypic manifestations of rare single-gene genetic disorders. Current measures of tissue specificity are unstable as they fail to account for similarity in the sample set used to define the transcriptome. Here, we introduce a novel method to measure tissue specificity that accounts for this similarity. To test measure robustness, brain subregion samples were successively added to a whole-body transcriptome sample set. As the proportion of brain subregion samples in the transcriptome increased from 1/42 samples to 13/54 samples, accounting for sample similarity resulted in 64% less variance in the change in specificity scores on average than when sample similarity was not considered. Ultimately, this method facilitates a more robust and reproducible measure of tissue specificity of gene expression.
WADE: Exploring the impact of recessive lethal mutations on inference of the distribution of fitness effects using genetic variation data
EMMA WADE1, Chris Kyriazis2, Kirk Lohmueller2,3
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Ecology and Evolutionary Biology, UCLA
3 Department of Human Genetics, David Geffen School of Medicine, UCLA
Whole genome sequencing has enabled inference of the DFE (distribution of fitness effects) from genetic variation data in large samples of individuals. However, some question the capability of these methods to detect the presence of lethal mutations. Here, we investigate SFS-based DFE inference methods and examine their potential to detect low frequency, recessive lethal mutations. Using simulation software, SLiM, we performed forward-in-time simulations with varying proportions of lethal alleles. Then, using our resulting SFSs and Python package Fitdadi, we fit and compared the inferred DFEs at each lethal proportion level. Although lethal mutations were present in our SFSs, we did not find a significant difference between the DFEs at varying proportions. This suggests DFE inference methods based on genetic variation data are indeed under-powered for detecting recessive lethal mutations. Future work is needed to improve our ability to quantify the extent of recessive lethal variation in humans and other species.
WANG J: A functional genomic alignment of human and mouse genomic regions
JEREMY WANG1, Soo Bin Kwon2,3, Runjia Li2,3, Jason Ernst2,3
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Bioinformatics Interdepartmental Program, UCLA
3 Department of Biological Chemistry, David Geffen School of Medicine, UCLA
Functional genomic correspondences between both sequence-aligned and non-sequence-aligned regions of the human and mouse genomes remain unclear. Since mice are established model organisms for understanding human disease, resolving this uncertainty is critical. Therefore, we develop ALignment by Learning Of an Immersion (ALLOI), which embeds human and mouse genomic regions in a common space based on their functional genomic profiles, allowing for geometric analyses of functional genomic similarity. ALLOI uses locally linear mappings in latent space, previously applied in single-cell data integration. By analyzing nearest neighbors in this space, we leverage characterized human chromatin states to study unknown mouse chromatin states. Further, despite the generally well-mixed distribution of the two species throughout ALLOI space, we identify clusters of genomic regions exclusively from one species and demonstrate that these regions have functional differences between humans and mice. Together, these results suggest that ALLOI is useful for transporting genomic understanding between these species.
WANG A: Machine Learning for the Prediction of COVID-19 Severity Levels From Salivaomics Data
AARON WANG1, Fang Wei2, David Wong2
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 School of Dentistry and Department of Pathology, UCLA
The clinical spectrum of COVID-19 is broad, extending from asymptomatic to severe immuno-pulmonary reactions that, if not categorized properly, may be life-threatening. Clinicians rate COVID-19 patients on a scale from 1 to 6 demonstrating the severity level of the patient, 1 being healthy and 6 being extremely sick, based on a multitude of clinical factors. However, there exists two issues with this severity level designation. Firstly, there exists variation among clinicians in presenting these patient scores, which may lead to improper treatment. Secondly, clinicians use a variety of metrics to determine patient severity level, including metrics involving plasma collection that require invasive procedures. This project attempts to alleviate both issues by introducing an ML framework that unifies severity level designations based on noninvasive saliva biomarkers. Our results show that we can successfully use ML on salivaomics data to predict COVID-19 severity, indicating the presence of viral load in saliva biomarkers.
YAKUBU, ZAKI: Analyzing cell populations and differentiation pathways in developing E12.5 mice mandibles
ABDUL-RAKEEM YAKUBU1, RAAFAE ZAKI1, Leah Ye2, Jimmy Hu2
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 School of Dentistry, University of California Los Angeles, Los Angeles, CA, USA
Craniofacial tissue provides models for studying progenitor cell specification and differentiation during mouse embryonic development. The formation of ectodermal appendages, such as the tooth, within the mouse mandible allow for understanding the genetic and signaling regulation of different cell subpopulations during tissue morphogenesis. However, questions remain on the cellular heterogeneity in the developing mandible and the regulatory interactions between different cell types. Utilizing single cell RNA-sequencing, this project strives to unveil gene expression patterns in order to map cell clusters to known cell types while inferring future differentiation pathways. Using the several tools within R, we analyzed cell clustering and differentiation patterns in embryonic day (E) 12 mouse mandible tissue to visualize differential gene expression and predict the trajectory of progenitor cell differentiation during embryonic development. Various tooth and mesenchymal populations were identified on the UMAP plot based on differentially expressed marker genes. Additional clusters of vasculature and muscle cells were also identified and separated from tooth and other mesenchymal cells. Early progenitors were identified in the molar regions towards the back of the mandible, where was predicted to differentiate into cartilage, incisor, and molar cells over the course of development. Identifying specific differentiation pathways can provide greater insight into the complex mechanisms underlying tissue formation in developing embryos. Such insight can be applied to future stem cell studies and regenerative medicine therapies.
YANG: Virus-host stability and coexistence in a modified predator-prey model
NICKIE YANG1, Chibundu Umunna1, Benjamin Knowles2
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Ecology and Evolutionary Biology, UCLA
While viruses and hosts coexist in nature, host populations generally collapse in theoretical models. Reconciling this inconsistency is a central question in viral ecology. Viruses can be lytic predators or temperate parasites, where they replicate within their hosts. We investigated what conditions favored temperate coexistence vs. lytic collapse by allowing the viruses to ‘choose’ whether to be lytic or not in each generation by optimizing host and viral population sizes. This resulted in stability and coexistence of both populations at low nutrient concentrations. By contrast, viruses collapsed hosts at higher nutrient concentrations, where infection rates were – impossibly – greater than host population size. We therefore made the infection rate equal to the smallest host or virus population. This led to a remarkably stable virus-host coexistence conserved across changing nutrient availability, as observed in nature. Together, our models captured natural virus-host stability and coexistence, parsimoniously reconciling theoretical models with empirical observation.
ZHA: Examining Effect of PGC-1Α Induction in the Mouse Oral Microbiome
LEON ZHA1, Troy Sisson2, Bhumika Shokeen3, Marcia Dinis3, Nini C. Tran4, Bo Yu5, Renate Lux3
1 BIG Summer Program, Institute for Quantitative and Computational Biosciences, UCLA
2 Department of Chemistry and Biochemistry, UCLA
3 Section of Periodontics, School of Dentistry, UCLA
4 Section of Pediatric Dentistry, School of Dentistry, UCLA
5 Section of Rest. Dentistry, School of Dentistry, UCLA
Periodontitis is an oral microbial disease that can damage gums and bone. Previous studies have shown resveratrol nanoparticles (RSV) attenuate periodontitis via PGC-1α induction and protect against alveolar bone loss. Here, we investigate the effect of RSV on the oral microbiome. Mice were inoculated with Porphyromonas gingivalis and Fusobacterium nucleatum to induce periodontitis, as well as RSV or no RSV. Oral samples were taken pre-inoculation, immediately after inoculation, and 1 month post-inoculation, processed for sequencing of the V1-V3 region of the 16S rRNA gene, and analyzed using QIIME2. We discovered that the mice receiving RSV treatment had increased species richness and abundance compared to their initial pre-inoculation state. On the other hand, the diversity of the oral microbiome in mice without RSV decreased after inoculation. These results suggest that RSV may provide a twofold protection against periodontitis by targeting host factors and influencing microbiome composition directly or via the effect on the host response.

